SRIBD News
Dr. Haofeng Li's Team From the Medical Big Data Laboratory Won the Runner-up in the NeurIPS 2022 CellSeg Competition
Recently, Dr. Li Haofeng's team from the Medical Big Data Laboratory of Shenzhen Research Institute of Big Data won the runner-up in the CellSeg competition of NeurIPS 2022, a top machine learning conference. The full name of the CellSeg competition is Weakly Supervised Cell Segmentation in Multi-modality High-Resolution Microscopy Images. Jointly organized by Broad Institute of the United States and other institutions, this competition attracted more than 100 teams from the Korean Academy of Science and Technology, the Jülich Research Center in Germany, Zhejiang University, and Nanjing University. Dr. Haofeng Li's team, including Lou Wei, a joint PhD student of the research institute, Xinyi Yu, a full-time engineer, and Chenyu Liu, an intern, represented the Shenzhen Big Data Research Institute under the team name SribdMed and won the runner-up.
Competition official website:
https://neurips22-cellseg.grand-challenge.org/awards/
Cell segmentation is often the first step in single-cell analysis tasks in microscope image-based biological and biomedical research. Deep learning models have been widely used in image segmentation. However, because manually labeling images is extremely time-consuming and expensive, it is difficult to collect a large number of labeled cell images to train the model. In addition, the datasets are usually limited to one modality and lack diversity, resulting in poor generalization ability of deep learning models. This competition constructed a multi-modal semi-supervised cell segmentation dataset, requiring participants to construct a general cell segmentation algorithm that can accurately segment cells of various modalities. At the same time, the algorithm is also required to be efficient enough to meet the running time requirement when segmenting large microscopic images.
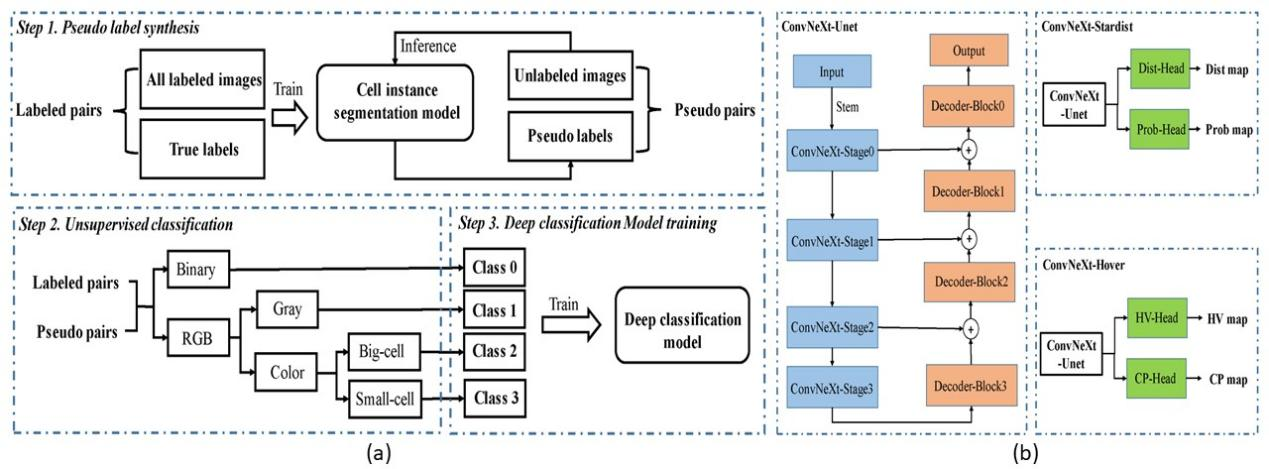
In the final solution, our team first built an automated classification model (shown in (a) above), which classifies cell images into four categories according to the low-level feature attributes of the images. Secondly, according to the morphological characteristics of different types of cells, we combined two existing segmentation models Stardist and HoverNet to learn to segment different types of cell images (as shown in (b) above). In order to build an efficient segmentation algorithm, the research team also deployed a high-efficiency neural network encoder -- ConvNeXt, making the final scheme tied for first place in terms of running time. In terms of segmentation accuracy and running time, this scheme finally won the runner-up.
Paper:
https://openreview.net/forum?id=G24BybwKe9
Code:
https://github.com/lhaof/CellSeg



